There has been enormous progress in our understanding of the complexity and diversity of the human genome, starting with the discovery of the structure of DNA1 and accelerating with the completion of the Human Genome Project in 2003.2 This has aided gene discovery for Mendelian disorders and progress towards the goal of ‘precision medicine’.3 In this article, we will focus on the application of genomics to the diagnosis of Mendelian genetic disorders and resulting clinical implications.
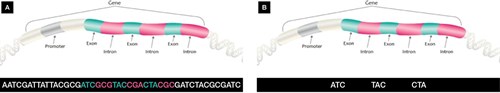
Our DNA or genomic sequence is made up of protein-coding amino acid sequences called genes, as well as large non-coding regions, where many regulatory elements are located. Genes consist of alternating regions of coding sequence called exons and non-coding sequences called introns (Figure 1).
Next generation sequencing
Figure 1. Difference between whole-genome sequencing and whole-exome sequencing
A. Whole-genome sequencing examines the entire DNA sequence that makes up the genome; B. Whole-exome sequencing sequences only the protein-coding regions (exons) of the genome, which account for only 1–2% of the whole genome but contain at least 85% of known disease-causing mutations.
Adapted from Genomics Education, www.genomicseducation.hee.nhs.uk
Initially, the process of determining the order of nucleotides within the human genome (known as sequencing) relied on techniques that were labour-intensive and cost-intensive. These barriers were overcome by the development of a high-throughput sequencing platform termed next-generation sequencing (NGS) or massively parallel sequencing. The three types of NGS that are currently being used in clinical practice are gene panel testing, whole-exome sequencing and whole-genome sequencing.
NGS has not superseded all genetic testing. Traditional cytogenetic and molecular testing methods are still indicated for specific conditions (Table 1). These tests include Sanger sequencing, which compares the amino acid sequence of an individual gene to a reference gene sequence to identify a pathogenic variant (mutation). Other commonly used testing methods are chromosome analysis, polymerase chain reaction (PCR) and DNA methylation studies.
Genetic testing is performed throughout Australia in private and public laboratories. Specific tests for rare conditions tend to be performed by one or a few preferred laboratories that have experience with the disease type or genes of interest. A service directory for these tests and laboratories is provided by the Royal College of Pathologists of Australasia (Table 2). Genetic tests for rare disorders or those not publically funded are requested by clinical geneticists.
Table 1. Types of genetic tests
|
Genetic test
|
Indication
|
Clinical example
|
|---|
|
G banded (conventional) karyotype
|
Suspected chromosome rearrangement
|
Multiple miscarriages or infertility
|
|
Fluorescent in situ hybridisation (FISH)
|
To determine physical arrangement of chromosomal abnormalities
|
Informing recurrence risk for parents of child with relevant chromosomal abnormality
|
|
Molecular karyotype (eg single-nucleotide polymorphism [SNP] microarray)
|
Submicroscopic chromosomal copy number variants
|
Intellectual disability
|
|
Sanger sequencing
|
Genetically homogeneous disorder
|
Neurofibromatosis type 2
|
|
When disorder typically results from 1–2 recurrent mutations, additional testing methods may be used for efficiency (eg PCR, geneprobe)
|
HFE-associated hereditary haemochromatosis
HLA alleles associated with coeliac disease
|
|
Polymerase chain reaction (PCR)
|
Disorder caused by intragenic triplet repeat expansions
|
Fragile X syndrome
Huntington’s disease
|
|
Multiplex ligation-dependent probe amplification (MLPA)
|
Disorder caused by multi nucleotide intragenic deletion
|
Duchenne muscular dystrophy
|
|
DNA methylation studies of specific chromosome region
|
Disorder caused by abnormal gene methylation
|
Beckwith–Wiedemann syndrome
|
|
Maternal serum screening/first trimester combined screening
|
Pregnancy at 11–13 weeks’ gestation to estimate risk for trisomy 21 or trisomy 18, neural tube defect or anencephaly
|
Low-risk pregnancy
|
|
Non-invasive prenatal test (NIPT), which tests cell free DNA in maternal blood (eg Percept, Harmony)
|
Pregnancy from 10 weeks onwards to detect evidence of trisomy 21, 18 or 13 in fetus with higher sensitivity and specificity than the maternal serum screening test
|
Advanced maternal age
|
Limitations of NGS testing
Limitations to the sensitivity and specificity of NGS as a diagnostic test currently exist because of technological challenges and knowledge gaps in understanding the significance of the results.
Technical challenges to accurate sequence detection largely relate to the presence of repetitive genomic arrangements that prevent short fragments of DNA aligning accurately to the sequencing platform. This is a barrier to the detection of expanded alleles in trinucleotide repeat disorders, as well as the sequencing of genes that are flanked by non-functional copies or pseudogenes, as is the case for CYP21A2-related congenital adrenal hyperplasia, CYP21A2 being closely flanked by the pseudogene CYP21A2P. In such situations, troubleshooting sequencing methods are used.
Gaps in our knowledge of benign and disease-causing genomic variations limit the accuracy with which we can assign clinical significance to an identified variant. The majority of the single nucleotide variants identified in a genome are polymorphisms, occurring in >1% of the population with no known impact on the function of the encoded protein.4 Features used to assess the clinical significance of a variant include its presence in population or disease-variant databases, or the medical literature. If novel, methods to predict the impact on protein translation and function are used to assess significance, along with testing of affected and unaffected family members to confirm segregation of the variant with disease.5 All of these methods provide evidence for or against the likelihood that a genetic variation is pathogenic. Universal genomic databanks and cell/animal model studies of gene function are necessary to improve our understanding of genomic variation. Figure 2 outlines an example of the exome data filtering process.
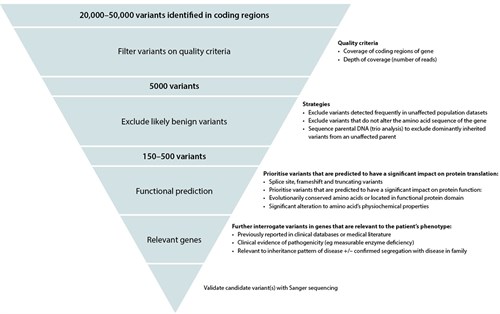
Figure 2. An example of a variant filtering process for exome sequencing data
Clinical applications
The first report using NGS to identify candidate genes for a small number of Mendelian-inherited conditions was published in 2009.6 NGS has since been applied to numerous cohorts of patients with unclassified genetic disease and this has been paralleled by a rapid rate of gene discovery. As an example, over 700 genes have now been identified in association with intellectual disability, and NGS contributed significantly to this advancement.7
The following case examples contrast the different approaches to, and applications of, genomic testing.
Gene panel testing
A gene panel uses NGS technology to simultaneously sequence a specific group of genes that are known to have mutations associated with a particular disease or clinical phenotype. Given that only the genes relevant to the disease are studied, the specificity of the test is increased. This feature makes the test more suitable for predictive testing of asymptomatic relatives, the application that currently yields the most benefit to clinical care. The clinical application of this test is demonstrated in Case 1.
Case 1. Hypertrophic cardiomyopathy
A healthy man, 30 years of age, with a family history of hypertrophic cardiomyopathy (HCM) is identified on screening echocardiography to have mild HCM. His mother died at the age of 52 years from this condition. This patient is the eldest of four siblings, all of whom have had a cardiac assessment since their mother’s diagnosis. His sister has mild HCM and his two brothers have normal echocardiograms. The genomic test applied in this case has a high specificity for cardiomyopathy-associated pathogenic variants. The patient’s cardiomyopathy panel identifies a pathogenic missense mutation in MYH7. Familial testing identifies that his sister and one brother are mutation-positive. As disease penetrance is age-dependent, frequent cardiological evaluation for HCM and risk factors for sudden cardiac death for asymptomatic mutation carriers is justified until advanced age. The molecular diagnosis in this case helped inform the need for lifelong surveillance for pre-symptomatic family members.
Whole-exome sequencing
Whole exome sequencing is the simultaneous sequencing of all of the exons, the protein-coding regions of the genome. This region makes up only 1–2% of the human genome but contain at least 85% of known disease-causing mutations.5 Analysing this small portion of the genome greatly reduces the amount of information to be filtered out and the cost of the process. A ‘clinical exome’ sequences only those genes that have a known association with disease, which reduces the exome from approximately 22,000 genes to approximately 3,000.
The diagnostic yield from NGS varies according to the disease being investigated and the stage in the diagnostic process at which NGS is applied. When applied to phenotypically diverse patient cohorts, the diagnostic yield from NGS can be as low as 25%.8 By contrast, large cohort studies that have applied NGS to the diagnosis of intellectual disability have achieved a diagnosis in 35%7 and demonstrated that 60% of the causative genetic mutations in intellectual disability arise de novo and are not inherited.9 Depending on whether it is used as a first-line investigation or a last resort, the yield can range from 57.7%10 to 45%11 respectively. The diagnostic yield of NGS is increasing as more disease-causing genes are identified and the phenotypic and mutational spectrum associated with known genes is better understood. This is illustrated further in Case 2.
Case 2. The child with developmental delay
A boy, eight months of age, is referred to a paediatric genetic service because of failure to thrive, microcephaly, hypoplastic thumbs, global developmental delay, and repaired trachea-oesophageal fistula. The differential diagnosis includes genetic causes for microcephaly, syndromes associated with radial ray defects, chromosome breakage disorders and DNA-repair defects. A molecular diagnosis in this case will inform prognosis, in particular the risk of bone marrow failure and malignancy, and need for surveillance. It will also inform reproductive risk for the family and enable prenatal diagnosis in future pregnancies. As there are several differential diagnoses for this case and a large number of candidate genes, single-gene sequencing would be costly, time-consuming and potentially non-diagnostic because of the genetic heterogeneity of this child’s phenotype. The most efficient diagnostic test in this setting is whole-exome sequencing with targeted analysis of a candidate gene list.10
In this case, whole-exome sequencing identified two previously reported pathogenic variants in the gene FANCA (Fanconi anaemia, complementation group A), one inherited from each parent, and the boy was diagnosed with autosomal recessive Fanconi anaemia. The child was referred for annual evaluation by a multidisciplinary team to monitor for complications of this disorder. As there will be a 25% recurrence risk for future pregnancies, the child’s parents were referred to a prenatal genetics clinic to discuss the options of prenatal testing and pre-implantation genetic testing that are now available to them if they choose.
Genome-wide association studies
Genome-wide association studies (GWAS) identify genetic variants that increase risk for common complex, multifactorial disorders, such as cardiovascular disease, diabetes mellitus and schizophrenia. This analysis compares the frequencies of genome-wide single-nucleotide polymorphisms (SNPs) in populations of affected and unaffected individuals.12 This research contributes to our understanding of disease pathogenesis, phenotypic variability and individual responses to certain drugs but, at present, rarely has an impact on clinical practice.13
Implications for general practice
The translation of genomic medicine into clinical care is increasing the opportunity for accurate genetic diagnosis and improved patient care. Access to genomic testing can avoid the traumatic ‘diagnostic odyssey’ for patients and their families and, in some cases, a timely diagnosis can change management. Diagnoses also enable prenatal testing, and may lead to the identification of at-risk family members where early treatment may significantly alter their outcome.10 Although general practitioners (GPs) may not be initiating NGS testing in many cases, genetic diagnoses identified in their patients may have an impact on their primary care management and identify risks to family members and future offspring.
Referral to a clinical genetics service may be useful when a genetic aetiology is suspected, such as a familial cancer, cardiomyopathy or hypercholesterolaemia, or for a child with severe and/or multiple congenital or developmental abnormalities, and in families concerned about recurrence risk for disease. Referral in such cases is often made by the oncologist, cardiologist or paediatrician involved but can be initiated by a GP. Referral is also indicated if support is needed for the diagnosis and/or management of a genetic disease. A comprehensive list of referral services for each state is available at the NSW government Centre for Genetics Education (www.genetics.edu.au/Genetics-Services).
With the expanding opportunities for genetic testing, selecting the appropriate genetic testing for patients becomes more difficult. Given that many genetic tests are not subsidised through the Medicare Benefits Scheme, and usually cost hundreds to thousands of dollars, it is essential to consider the likely diagnostic yield of a test in each circumstance; a clinical genetics service can assist with this process. Although there can be long waiting times to access a clinical genetics service, support from a genetics specialist regarding test selection and interpretation of results may save significant time and money in the long term. Education and support provided by a clinical genetics service can also help patients and clinicians maximise the benefits of genomic testing while reducing potential harms.14 In the consent process, patients can also agree to the global sharing of their NGS data, facilitating the genetic research that is an integral component of clinical genetics practice.
Conclusion
Genomic testing is becoming increasingly available to patients and clinicians, and rapidly being integrated into routine clinical care. There are many advantages to this technology, but also limitations, and it is essential that GPs consider these in their management of patients and families affected by genetic disease.
Authors
Lilian Downie MBBS (Hons), Clinical Genetics Fellow, Murdoch Childrens Research Institute, Victorian Clinical Genetics Services, Royal Children’s Hospital, Parkville, Vic, University of Melbourne, Vic
Sarah Donoghue BMedSci (Hons), MBBS (Hons), DCH, FRACP, Clinical Genetics Fellow, Murdoch Childrens Research Institute, Victorian Clinical Genetics Services, Royal Children’s Hospital, Parkville, Vic
Chloe Stutterd BHSc, MBBS, FRACP, Clinical Geneticist, Murdoch Childrens Research Institute, Victorian Clinical Genetics Services, Royal Children’s Hospital, Parkville, Vic; PhD candidate, University of Melbourne, Vic. chloe.stutterd@mcri.edu.au
Competing interests: None.
Provenance and peer review: Commissioned, externally peer reviewed